To view your price please login or contact us
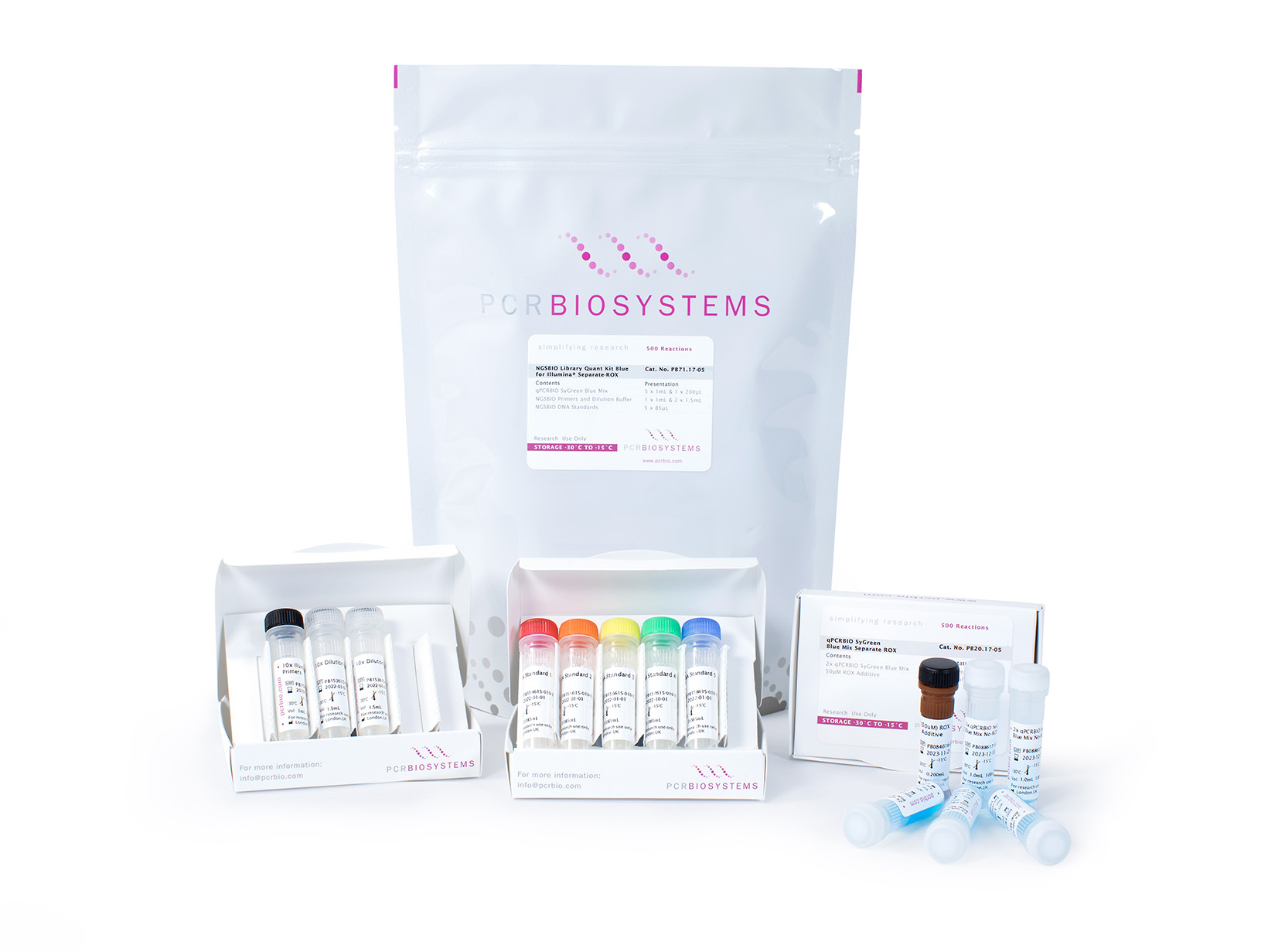
NGSBIO Library Quant Kit Blue for Illumina®
The NGSBIO Library Quant Kit Blue contains all the components required for accurate and sensitive quantification of libraries prepared for Illumina® NGS systems.
The kit uses an easy-to-see blue qPCR mix to specifically quantify adapter-ligated DNA molecules, ensuring optimal cluster densities for improved sequencing efficiency and quality of data.
Features
- Easy-to-see blue mix for greater pipetting precision and accuracy
- Uses qPCR to accurately and rapidly quantify a library prior to sequencing
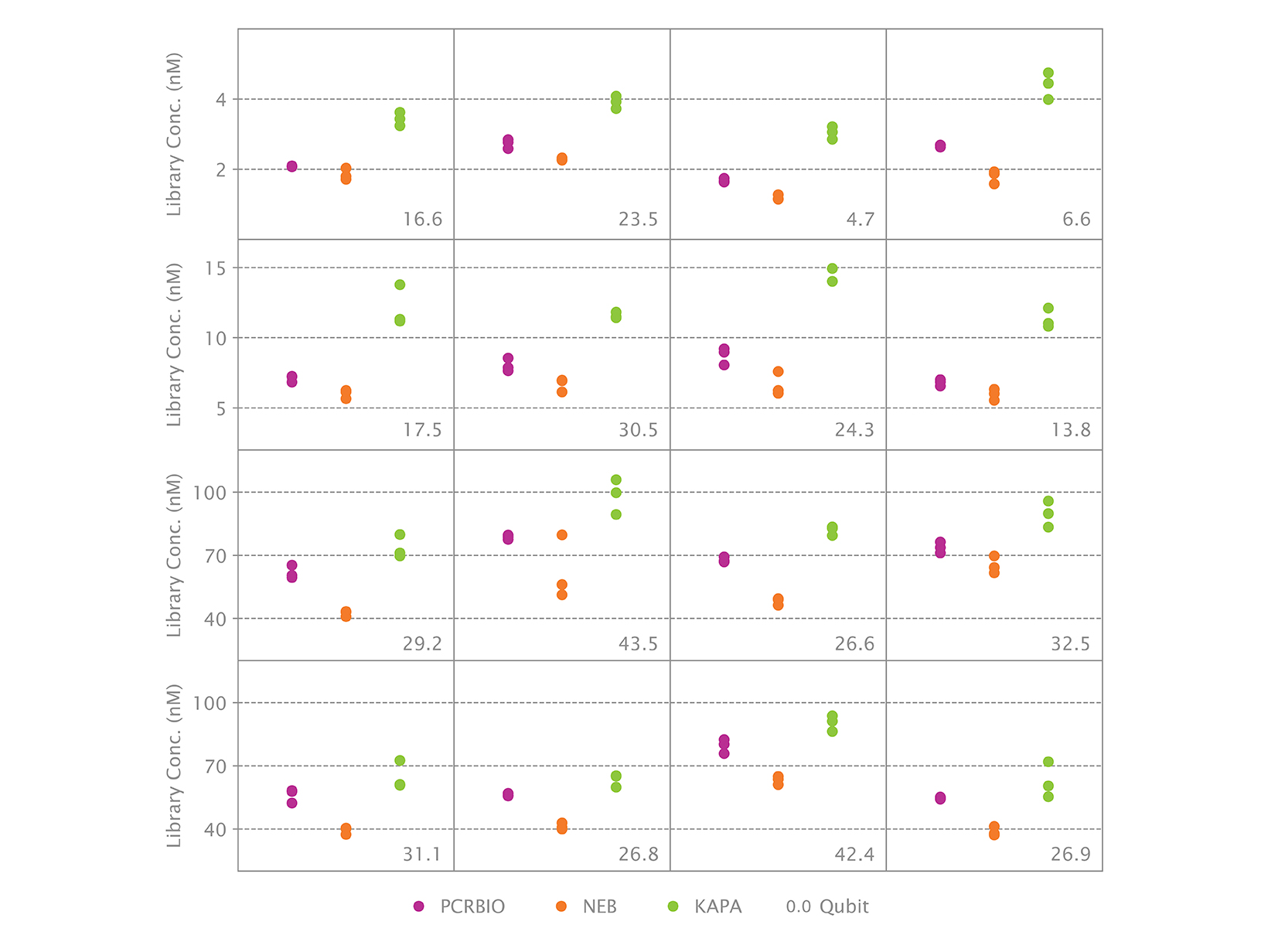
- Gives consistent library quantification across a wide range of sample types, concentrations, fragment sizes and GC content
- Uses a single extension time for all libraries
- Allows specific quantification of only DNA molecules that can be sequenced by NGS
- Uses antibody-mediated hot start technology to ensure all reactions start simultaneously
- Compatible with all Illumina® instruments and qPCR platforms
- Suitable for manual and automated workflows
- Easily calculate library concentration with the online NGSBIO Library Quantification Tool
More Information
The NGSBIO Library Quant Kit Blue offers a reliable qPCR-based method for the quantification of libraries prepared for Illumina® NGS systems. The kit includes 5 DNA standards, primers specific to the P5 and P7 Illumina® adapter sequences and qPCRBIO SyGreen Blue Mix. Our blue qPCR mixes contain a non-reactive dye to improve reaction mix visibility, allowing greater pipetting precision and reduced errors without affecting your real-time PCR performance. The advanced qPCR buffer system has been developed using our high-throughput smart screen technology to ensure efficient amplification of all your libraries, including those that are GC or AT-rich. The kit is also supplied with a convenient library dilution buffer and can also be used in combination with VeriFi™ Library Amplification Mix.
Accurate Library Quantification
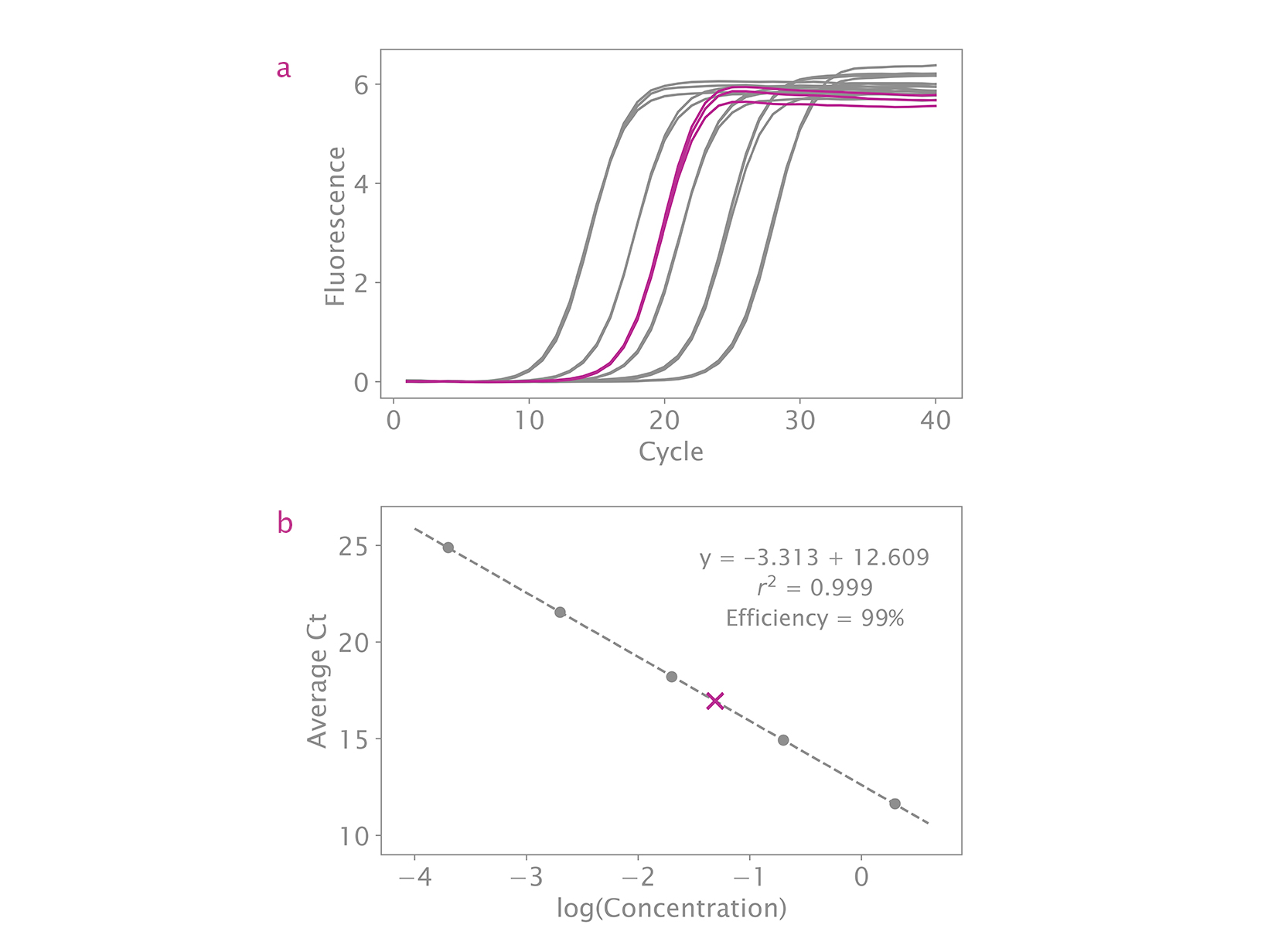
qPCR is considered the best method for quantifying NGS libraries as it only measures adapter-bound molecules that can be used as templates for library amplification and cluster generation. The NGSBIO Library Quant Kit enables highly accurate quantification crucial for optimal cluster densities and greater sequencing efficiency.
Library quantification can be used at any time after adapter ligation and should always be used prior to cluster generation. Overestimating library concentration can result in insufficient cluster density and underestimating library concentration can result in a high, saturating cluster density. To learn more about optimal cluster density, consult the instructions of your Illumina® machine.
Wide Dynamic Range
The DNA standards supplied are precisely measured and ready-to-use, covering 5 orders of magnitude from 2pM to 0.2fM. The kit is suitable for quantification of even low concentration libraries including libraries constructed without a PCR amplification step.
Universal Kit
The NGSBIO Library Quant Kit is compatible with all qPCR platforms and is optimised to give consistent and reproducible library quantification across a wide range of sample types, fragment sizes (up to 1000bp), concentrations and GC content. Use our qPCR Selection Tool to find out which ROX variant is compatible with your instrument.
For easy and accurate data analysis we recommend using the NGSBIO Library Quantification Tool.
Applications
- NGS library quantification for Illumina® systems
Specifications
NGSBIO Library Quant Kit Blue for Illumina® Lo-ROX
Component
100 Reactions
500 Reactions
2x qPCRBIO SyGreen Blue Mix Lo-ROX
1 x 1mL
5 x 1mL
DNA Standards 1-5
30μL each
85μL each
10x Illumina® Primers
1 x 0.2mL
1 x 1mL
10x Dilution Buffer
1 x 0.6mL
2 x 1.5mL
NGSBIO Library Quant Kit for Illumina® Hi-ROX
Component
100 Reactions
500 Reactions
2x qPCRBIO SyGreen Blue Mix Hi-ROX
1 x 1mL
5 x 1mL
DNA Standards 1-5
30μL each
85μL each
10x Illumina® Primers
1 x 0.2mL
1 x 1mL
10x Dilution Buffer
1 x 0.6mL
2 x 1.5mL
NGSBIO Library Quant Kit for Illumina® Separate-ROX
Component
100 Reactions
500 Reactions
2x qPCRBIO SyGreen Blue Mix No-ROX
1 x 1mL
5 x 1mL
50μM ROX Additive
1 x 200μL
1 x 200μL
DNA Standards 1-5
30μL each
85μL each
10x Illumina® Primers
1 x 0.2mL
1 x 1mL
10x Dilution Buffer
1 x 0.6mL
2 x 1.5mL
NGSBIO DNA Standards for Illumina®
Component
500 Reactions
DNA Standards 1-5
85μL each
NGSBIO Library Quant Kit Blue for Illumina® Lo-ROX
Component
2x qPCRBIO SyGreen Blue Mix Lo-ROX
DNA Standards 1-5
10x Illumina® Primers
10x Dilution Buffer
100 Reactions
1 x 1mL
30μL each
1 x 0.2mL
1 x 0.6mL
500 Reactions
5 x 1mL
85μL each
1 x 1mL
2 x 1.5mL
NGSBIO Library Quant Kit for Illumina® Hi-ROX
Component
2x qPCRBIO SyGreen Blue Mix Hi-ROX
DNA Standards 1-5
10x Illumina® Primers
10x Dilution Buffer
100 Reactions
1 x 1mL
30μL each
1 x 0.2mL
1 x 0.6mL
500 Reactions
5 x 1mL
85μL each
1 x 1mL
2 x 1.5mL
NGSBIO Library Quant Kit for Illumina® Separate-ROX
Component
2x qPCRBIO SyGreen Blue Mix No-ROX
50μM ROX Additive
DNA Standards 1-5
10x Illumina® Primers
10x Dilution Buffer
100 Reactions
1 x 1mL
1 x 200μL
30μL each
1 x 0.2mL
1 x 0.6mL
500 Reactions
5 x 1mL
1 x 200μL
85μL each
1 x 1mL
2 x 1.5mL
NGSBIO DNA Standards for Illumina®
Component
DNA Standards 1-5
500 Reactions
85μL each
Reaction Volume
Storage
20μL
On arrival, products should be stored between -30 and -15°C. If stored correctly the kit will retain full activity for 12 months.
Reaction Volume
20μL
Storage
On arrival, products should be stored between -30 and -15°C. If stored correctly the kit will retain full activity for 12 months.
Instrument Compatibility
This product is compatible with all standard and fast cycling qPCR instruments. Use our qPCR Selection Tool to find out which ROX variant is compatible with your instrument.
Documents
Product Flyers
Product Manuals
Material Safety Data Sheets
Certificate of Analysis Finder
FAQs
My samples fall outside of the range of standards. What troubleshooting suggestions do you have?
Examine the melt curve to ensure that there is a single peak. If you see multiple peaks, there may be contamination present or unligated adapters. Otherwise, if the samples appear before DNA Standard 1, dilute the sample 100x and repeat. If the samples appear after DNA Standard 5, use a sample that is 100x more concentrated.
My standard curves are not evenly spaced. Does this matter?
If the overall efficiency of your standard curve is between 90% – 110% and your R2 value is close to 1, this is not a cause for concern. As long as the slope of the line through all of the standards is between 3.1 – 3.6, the quantification will be accurate.
The efficiency of my standard curve is not within 90 – 110%. What troubleshooting suggestions do you have?
Inspect the curve for outliers. If any of the data points appear to be far from the fitted curve, delete that data point and see if the efficiency improves. A minimum of three standards is required for a reliable standard curve, so up to two standards can be deleted entirely. However, each standard needs at least two measurements, and only samples that fall within the remaining standards can be quantified.
Inspect the No Template Control (NTC) to ensure it has a greater Cq than DNA Standard 5. If the NTC falls within the standards’ range, there may be contamination present in the standards or reaction mix. This may come from other samples on the plate, or traces of other libraries remaining on laboratory surfaces or equipment. Clean all instruments and surfaces thoroughly and repeat the experiment with fresh aliquots of reagents.
What does it mean if I see other significant peaks in the melt analysis of my samples?
If the peak overlaps with the shoulder seen in the standards’ melting peaks, this is likely amplification of the Illumina® primers. This can occur when the primer/template ratio is too high, or when there are large amounts of unligated adapters. If the peak is significant, the quantification results are not reliable and the experiment should be repeated. Ensure that the correct amount of primer is added to the master mix and use a higher concentration of template if possible. If the problem persists, the library may need to go through size selection again to remove free adapters.
If multiple peaks are observed that do not correspond to the adapter peak, the most likely causes are either contamination or unsuccessful size selection during library preparation. Repeat the measurement with fresh dilutions of the library. If the problem persists, that library may need to undergo further size selection.
What does it mean if I see other significant peaks in the melt analysis of my standards?
It is normal to see a slight shoulder on the low-temperature side of the sample melt peak. If the shoulder is very large, either extra Illumina® Primer mix was added or an insufficient amount of standard was added. In this case, the experiment should be repeated with special care taken while making the master mix and when adding standards to the reaction.
If multiple peaks are observed in the melt curves of the standards, then contamination was introduced into the reaction. If it is present in all of the standards, the contamination is in the master mix. The experiment should be repeated with new aliquots of reagents. If it is present in the replicates of only one standard, that standard aliquot has become contaminated and must be discarded.
What if my R2 value is not between 0.99 and 1.00?
If the R2 value is not close to 1, this suggests that the standards are not spaced evenly. Inspect the standard curve chart and remove any points that are far from the line. As long as there are three standards with two measurements each, the quantification will be accurate.
If you are unable to bring the quality of the fit to the necessary level, the experiment should be repeated. Take extra care when pipetting the standards to ensure accurate volumes and prevent contamination.
What if the measurement I get with my NGSBIO Library Quant Kit does not agree with my Qubit, spectrophotometer, or another method measurement?
Instruments like the Qubit or a spectrophotometer measure the total amount of DNA present in solution. The same is true of concentration measurements by an Agilent Bioanalyser. The NGSBIO Library Quant Kit for Illumina® will only quantify molecules with both adapters present. In addition, qPCR is a much more accurate method in general for quantifying DNA compared to fluorescent or spectroscopic methods. It is normal for these different measurement methods to disagree.
If the concentration measured by qPCR is significantly lower than what is measured by another method, this suggests that only a small proportion of molecules correctly incorporated the adapters during adapter ligation.
What should I do if my sample replicates show significant variation?
Check the melt curves to ensure there is a single product peak. If there are multiple peaks, this suggests the sample may have become contaminated or degraded, or that an error occurred during the size selection or adapter ligation steps or library preparation. If the melt analysis shows only one peak, the variation is probably caused by inaccurate pipetting. Create a new dilution of the sample and repeat the experiment, taking care to pipette the replicates accurately. If the problem persists, a new library may need to be created.
What should I do if my standards replicates show significant variation?
A variation of ± 0.2 cycles between replicates is normal. Check the melt curves to ensure there is a single product peak. (There may be a lower molecular weight shoulder present, this is normal). If there are multiple peaks, this suggests the standards have become contaminated or degraded. If that standard can be removed from the analysis, do so. Otherwise, order new standards and repeat the experiment.
If the melt curves show a single peak that overlaps between replicates, the variation is probably caused by inaccurate pipetting. Repeat the experiment, taking care to pipette the standards accurately.
More Information
The NGSBIO Library Quant Kit Blue offers a reliable qPCR-based method for the quantification of libraries prepared for Illumina® NGS systems. The kit includes 5 DNA standards, primers specific to the P5 and P7 Illumina® adapter sequences and qPCRBIO SyGreen Blue Mix. Our blue qPCR mixes contain a non-reactive dye to improve reaction mix visibility, allowing greater pipetting precision and reduced errors without affecting your real-time PCR performance. The advanced qPCR buffer system has been developed using our high-throughput smart screen technology to ensure efficient amplification of all your libraries, including those that are GC or AT-rich. The kit is also supplied with a convenient library dilution buffer and can also be used in combination with VeriFi™ Library Amplification Mix.
Accurate Library Quantification
qPCR is considered the best method for quantifying NGS libraries as it only measures adapter-bound molecules that can be used as templates for library amplification and cluster generation. The NGSBIO Library Quant Kit enables highly accurate quantification crucial for optimal cluster densities and greater sequencing efficiency.
Library quantification can be used at any time after adapter ligation and should always be used prior to cluster generation. Overestimating library concentration can result in insufficient cluster density and underestimating library concentration can result in a high, saturating cluster density. To learn more about optimal cluster density, consult the instructions of your Illumina® machine.
Wide Dynamic Range
The DNA standards supplied are precisely measured and ready-to-use, covering 5 orders of magnitude from 2pM to 0.2fM. The kit is suitable for quantification of even low concentration libraries including libraries constructed without a PCR amplification step.
Universal Kit
The NGSBIO Library Quant Kit is compatible with all qPCR platforms and is optimised to give consistent and reproducible library quantification across a wide range of sample types, fragment sizes (up to 1000bp), concentrations and GC content. Use our qPCR Selection Tool to find out which ROX variant is compatible with your instrument.
For easy and accurate data analysis we recommend using the NGSBIO Library Quantification Tool.
Applications
- NGS library quantification for Illumina® systems
Specifications
NGSBIO Library Quant Kit Blue for Illumina® Lo-ROX
Component
100 Reactions
500 Reactions
2x qPCRBIO SyGreen Blue Mix Lo-ROX
1 x 1mL
5 x 1mL
DNA Standards 1-5
30μL each
85μL each
10x Illumina® Primers
1 x 0.2mL
1 x 1mL
10x Dilution Buffer
1 x 0.6mL
2 x 1.5mL
NGSBIO Library Quant Kit for Illumina® Hi-ROX
Component
100 Reactions
500 Reactions
2x qPCRBIO SyGreen Blue Mix Hi-ROX
1 x 1mL
5 x 1mL
DNA Standards 1-5
30μL each
85μL each
10x Illumina® Primers
1 x 0.2mL
1 x 1mL
10x Dilution Buffer
1 x 0.6mL
2 x 1.5mL
NGSBIO Library Quant Kit for Illumina® Separate-ROX
Component
100 Reactions
500 Reactions
2x qPCRBIO SyGreen Blue Mix No-ROX
1 x 1mL
5 x 1mL
50μM ROX Additive
1 x 200μL
1 x 200μL
DNA Standards 1-5
30μL each
85μL each
10x Illumina® Primers
1 x 0.2mL
1 x 1mL
10x Dilution Buffer
1 x 0.6mL
2 x 1.5mL
NGSBIO DNA Standards for Illumina®
Component
500 Reactions
DNA Standards 1-5
85μL each
NGSBIO Library Quant Kit Blue for Illumina® Lo-ROX
Component
2x qPCRBIO SyGreen Blue Mix Lo-ROX
DNA Standards 1-5
10x Illumina® Primers
10x Dilution Buffer
100 Reactions
1 x 1mL
30μL each
1 x 0.2mL
1 x 0.6mL
500 Reactions
5 x 1mL
85μL each
1 x 1mL
2 x 1.5mL
NGSBIO Library Quant Kit for Illumina® Hi-ROX
Component
2x qPCRBIO SyGreen Blue Mix Hi-ROX
DNA Standards 1-5
10x Illumina® Primers
10x Dilution Buffer
100 Reactions
1 x 1mL
30μL each
1 x 0.2mL
1 x 0.6mL
500 Reactions
5 x 1mL
85μL each
1 x 1mL
2 x 1.5mL
NGSBIO Library Quant Kit for Illumina® Separate-ROX
Component
2x qPCRBIO SyGreen Blue Mix No-ROX
50μM ROX Additive
DNA Standards 1-5
10x Illumina® Primers
10x Dilution Buffer
100 Reactions
1 x 1mL
1 x 200μL
30μL each
1 x 0.2mL
1 x 0.6mL
500 Reactions
5 x 1mL
1 x 200μL
85μL each
1 x 1mL
2 x 1.5mL
NGSBIO DNA Standards for Illumina®
Component
DNA Standards 1-5
500 Reactions
85μL each
Reaction Volume
Storage
20μL
On arrival, products should be stored between -30 and -15°C. If stored correctly the kit will retain full activity for 12 months.
Reaction Volume
20μL
Storage
On arrival, products should be stored between -30 and -15°C. If stored correctly the kit will retain full activity for 12 months.
Instrument Compatibility
This product is compatible with all standard and fast cycling qPCR instruments. Use our qPCR Selection Tool to find out which ROX variant is compatible with your instrument.
Documents
Product Flyers
Product Manuals
Material Safety Data Sheets
Certificate of Analysis Finder
FAQs
My samples fall outside of the range of standards. What troubleshooting suggestions do you have?
Examine the melt curve to ensure that there is a single peak. If you see multiple peaks, there may be contamination present or unligated adapters. Otherwise, if the samples appear before DNA Standard 1, dilute the sample 100x and repeat. If the samples appear after DNA Standard 5, use a sample that is 100x more concentrated.
My standard curves are not evenly spaced. Does this matter?
If the overall efficiency of your standard curve is between 90% – 110% and your R2 value is close to 1, this is not a cause for concern. As long as the slope of the line through all of the standards is between 3.1 – 3.6, the quantification will be accurate.
The efficiency of my standard curve is not within 90 – 110%. What troubleshooting suggestions do you have?
Inspect the curve for outliers. If any of the data points appear to be far from the fitted curve, delete that data point and see if the efficiency improves. A minimum of three standards is required for a reliable standard curve, so up to two standards can be deleted entirely. However, each standard needs at least two measurements, and only samples that fall within the remaining standards can be quantified.
Inspect the No Template Control (NTC) to ensure it has a greater Cq than DNA Standard 5. If the NTC falls within the standards’ range, there may be contamination present in the standards or reaction mix. This may come from other samples on the plate, or traces of other libraries remaining on laboratory surfaces or equipment. Clean all instruments and surfaces thoroughly and repeat the experiment with fresh aliquots of reagents.
What does it mean if I see other significant peaks in the melt analysis of my samples?
If the peak overlaps with the shoulder seen in the standards’ melting peaks, this is likely amplification of the Illumina® primers. This can occur when the primer/template ratio is too high, or when there are large amounts of unligated adapters. If the peak is significant, the quantification results are not reliable and the experiment should be repeated. Ensure that the correct amount of primer is added to the master mix and use a higher concentration of template if possible. If the problem persists, the library may need to go through size selection again to remove free adapters.
If multiple peaks are observed that do not correspond to the adapter peak, the most likely causes are either contamination or unsuccessful size selection during library preparation. Repeat the measurement with fresh dilutions of the library. If the problem persists, that library may need to undergo further size selection.
What does it mean if I see other significant peaks in the melt analysis of my standards?
It is normal to see a slight shoulder on the low-temperature side of the sample melt peak. If the shoulder is very large, either extra Illumina® Primer mix was added or an insufficient amount of standard was added. In this case, the experiment should be repeated with special care taken while making the master mix and when adding standards to the reaction.
If multiple peaks are observed in the melt curves of the standards, then contamination was introduced into the reaction. If it is present in all of the standards, the contamination is in the master mix. The experiment should be repeated with new aliquots of reagents. If it is present in the replicates of only one standard, that standard aliquot has become contaminated and must be discarded.
What if my R2 value is not between 0.99 and 1.00?
If the R2 value is not close to 1, this suggests that the standards are not spaced evenly. Inspect the standard curve chart and remove any points that are far from the line. As long as there are three standards with two measurements each, the quantification will be accurate.
If you are unable to bring the quality of the fit to the necessary level, the experiment should be repeated. Take extra care when pipetting the standards to ensure accurate volumes and prevent contamination.
What if the measurement I get with my NGSBIO Library Quant Kit does not agree with my Qubit, spectrophotometer, or another method measurement?
Instruments like the Qubit or a spectrophotometer measure the total amount of DNA present in solution. The same is true of concentration measurements by an Agilent Bioanalyser. The NGSBIO Library Quant Kit for Illumina® will only quantify molecules with both adapters present. In addition, qPCR is a much more accurate method in general for quantifying DNA compared to fluorescent or spectroscopic methods. It is normal for these different measurement methods to disagree.
If the concentration measured by qPCR is significantly lower than what is measured by another method, this suggests that only a small proportion of molecules correctly incorporated the adapters during adapter ligation.
What should I do if my sample replicates show significant variation?
Check the melt curves to ensure there is a single product peak. If there are multiple peaks, this suggests the sample may have become contaminated or degraded, or that an error occurred during the size selection or adapter ligation steps or library preparation. If the melt analysis shows only one peak, the variation is probably caused by inaccurate pipetting. Create a new dilution of the sample and repeat the experiment, taking care to pipette the replicates accurately. If the problem persists, a new library may need to be created.
What should I do if my standards replicates show significant variation?
A variation of ± 0.2 cycles between replicates is normal. Check the melt curves to ensure there is a single product peak. (There may be a lower molecular weight shoulder present, this is normal). If there are multiple peaks, this suggests the standards have become contaminated or degraded. If that standard can be removed from the analysis, do so. Otherwise, order new standards and repeat the experiment.
If the melt curves show a single peak that overlaps between replicates, the variation is probably caused by inaccurate pipetting. Repeat the experiment, taking care to pipette the standards accurately.